Abstract
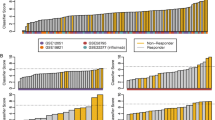
Our aim was to identify differentially expressed (DE) genes and biological processes that may help predict patient response to biologic agents for rheumatoid arthritis (RA). Using the INMEX (integrative meta-analysis of expression data) software tool, we performed a meta-analysis of publicly available microarray Gene Expression Omnibus (GEO) datasets that examined patient response to biologic therapy for RA. Three GEO datasets, containing 79 responders and 34 non-responders, were included in the meta-analysis. We identified 1,374 genes that were consistently differentially expressed in responders vs. non-responders (651 up-regulated and 723 down-regulated). The up-regulated gene with the smallest p value (p = 0.000192) was ASCC2 (Activating Signal Cointegrator 1 Complex Subunit 2), and the up-regulated gene with the largest fold change (average log fold change = −0.75869, p = 0.000206) was KLRC3 (Killer Cell Lectin-Like Receptor Subfamily C, Member 3). The down-regulated gene with the smallest p value (p = 0.000195) was MPL (Myeloproliferative Leukemia Virus Oncogene). Among the 236 GO terms associated with the set of DE genes, the most significantly enriched was “CTP biosynthetic process” (GO:0006241; p = 0.000454). Our meta-analysis identified genes that were consistently DE in responders vs. non-responders, as well as biological pathways associated with this set of genes. These results provide insight into the molecular mechanisms underlying responsiveness to biologic therapy for RA.


Similar content being viewed by others
References
Harris ED Jr (1990) Rheumatoid arthritis. Pathophysiology and implications for therapy. N Engl J Med 322(18):1277–1289
Brennan FM, Maini RN, Feldmann M (1992) TNF alpha—a pivotal role in rheumatoid arthritis? Br J Rheumatol 31(5):293–298
Lee YH, Woo JH, Rho YH, Choi SJ, Ji JD, Song GG (2008) Meta-analysis of the combination of TNF inhibitors plus MTX compared to MTX monotherapy, and the adjusted indirect comparison of TNF inhibitors in patients suffering from active rheumatoid arthritis. Rheumatol Int 28(6):553–559
Reff ME, Carner K, Chambers KS, Chinn PC, Leonard JE, Raab R et al (1994) Depletion of B cells in vivo by a chimeric mouse human monoclonal antibody to CD20. Blood 83(2):435–445
Clynes RA, Towers TL, Presta LG, Ravetch JV (2000) Inhibitory Fc receptors modulate in vivo cytoxicity against tumor targets. Nat Med 6(4):443–446
Golub TR, Slonim DK, Tamayo P, Huard C, Gaasenbeek M, Mesirov JP et al (1999) Molecular classification of cancer: class discovery and class prediction by gene expression monitoring. Science 286(5439):531–537
Choi SJ, Rho YH, Ji JD, Song GG, Lee YH (2006) Genome scan meta-analysis of rheumatoid arthritis. Rheumatology (Oxford) 45(2):166–170
Smith SL, Plant D, Eyre S, Barton A (2013) The potential use of expression profiling: implications for predicting treatment response in rheumatoid arthritis. Ann Rheum Dis 72(7):1118–1124
Ramasamy A, Mondry A, Holmes CC, Altman DG (2008) Key issues in conducting a meta-analysis of gene expression microarray datasets. PLoS Med 5(9):e184
Siddiqui AS, Delaney AD, Schnerch A, Griffith OL, Jones SJ, Marra MA (2006) Sequence biases in large scale gene expression profiling data. Nucleic Acids Res 34(12):e83
Cahan P, Rovegno F, Mooney D, Newman JC, St Laurent G 3rd, McCaffrey TA (2007) Meta-analysis of microarray results: challenges, opportunities, and recommendations for standardization. Gene 401(1–2):12–18
Rung J, Brazma A (2013) Reuse of public genome-wide gene expression data. Nat Rev Genet 14(2):89–99
Griffith OL, Melck A, Jones SJ, Wiseman SM (2006) Meta-analysis and meta-review of thyroid cancer gene expression profiling studies identifies important diagnostic biomarkers. J Clin Oncol 24(31):5043–5051
Lee YH, Rho YH, Choi SJ, Ji JD, Song GG (2005) Ankylosing spondylitis susceptibility loci defined by genome-search meta-analysis. J Hum Genet 50(9):453–459
Lee YH, Rho YH, Choi SJ, Ji JD, Song GG (2006) Meta-analysis of genome-wide linkage studies for bone mineral density. J Hum Genet 51(5):480–486
Xia J, Fjell CD, Mayer ML, Pena OM, Wishart DS, Hancock RE (2013) INMEX—a web-based tool for integrative meta-analysis of expression data. Nucleic Acids Res 41(W1):W63–W70
Barrett T, Troup DB, Wilhite SE, Ledoux P, Evangelista C, Kim IF et al (2011) NCBI GEO: archive for functional genomics data sets—10 years on. Nucleic Acids Res 39(Database issue):D1005–D1010
Arnett FC, Edworthy SM, Bloch DA, McShane DJ, Fries JF, Cooper NS et al (1988) The American Rheumatism Association 1987 revised criteria for the classification of rheumatoid arthritis. Arthritis Rheum 31(3):315–324
Bolstad BM, Irizarry RA, Astrand M, Speed TP (2003) A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19(2):185–193
Hong F, Breitling R, McEntee CW, Wittner BS, Nemhauser JL, Chory J (2006) RankProd: a bioconductor package for detecting differentially expressed genes in meta-analysis. Bioinformatics 22(22):2825–2827
Xia J, Fjell CD, Mayer ML, Pena OM, Wishart DS, Hancock RE (2013) INMEX—a web-based tool for integrative meta-analysis of expression data. Nucleic Acids Res. 41(Web Server issue):W63-70
Falcon S, Gentleman R (2007) Using GOstats to test gene lists for GO term association. Bioinformatics 23(2):257–258
Jung DJ, Sung HS, Goo YW, Lee HM, Park OK, Jung SY et al (2002) Novel transcription coactivator complex containing activating signal cointegrator 1. Mol Cell Biol 22(14):5203–5211
Lazetic S, Chang C, Houchins JP, Lanier LL, Phillips JH (1996) Human natural killer cell receptors involved in MHC class I recognition are disulfide-linked heterodimers of CD94 and NKG2 subunits. J Immunol 157(11):4741–4745
Matthews EE, Thevenin D, Rogers JM, Gotow L, Lira PD, Reiter LA et al (2011) Thrombopoietin receptor activation: transmembrane helix dimerization, rotation, and allosteric modulation. FASEB J 25(7):2234–2244
Jones C, Simpson P, Mackay A, Lakhani SR (2006) Expression profiling using cDNA microarrays. Methods Mol Med 120:403–414
Desvignes T, Pontarotti P, Fauvel C, Bobe J (2009) Nme protein family evolutionary history, a vertebrate perspective. BMC Evol Biol 9:256
Klein RJ, Zeiss C, Chew EY, Tsai JY, Sackler RS, Haynes C et al (2005) Complement factor H polymorphism in age-related macular degeneration. Science 308(5720):385–389
Acknowledgements
This study was supported in part by a grant of the Korea Healthcare technology R&D project, Ministry for Health & Welfare, Republic of Korea (HI12C1834)
Conflict of interest statement
The authors have no conflict of interest to declare.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(XLSX 27 kb)
Rights and permissions
About this article
Cite this article
Lee, Y.H., Bae, SC. & Song, G.G. Meta-analysis of gene expression profiles to predict response to biologic agents in rheumatoid arthritis. Clin Rheumatol 33, 775–782 (2014). https://doi.org/10.1007/s10067-014-2547-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10067-014-2547-9